Retinal dystrophies and optic neuropathies are the thrust areas of research. We utilise next-generation sequencing (NGS) to uncover the genetic factors associated with disease pathogenesis. Our approach offers accurate molecular diagnosis, thereby facilitating access to precise novel therapeutic interventions. Personalised genetic counselling was provided to the affected families. Our endeavour aims to construct a gene registry of the visual system specific to the South Indian ethnicity.
Molecular Biology:
Our lab is primarily focusing on translational eye research. We developed novel and affordable genetic testing methods for retinoblastoma, which is currently being used for routine clinical care. We extensively utilise multi-omic technologies to unravel the molecular mysteries in eye cancers. Different model systems, including drug resistant cell lines, patient-derived spheroids, and zebra fish models, are being deployed to test the drugs or small molecules for improving the treatment.
Currently, our lab is focused on characterising the genetic factors underlying rare neural degenerative disorders in the human visual system, such as Leber’s hereditary optic neuropathy and Leber’s congenital amaurosis.
Leber’s Congenital Amaurosis (LCA):
We identify additional LCA candidate genes/loci from existing study subjects negative for known LCA candidate genes through whole genome sequencing and genome wide association studies. This approach will help to develop an ethnic specific gene panel for molecular diagnosis. The biological relevance of the identified candidate genes will be functionally validated through in vitro experiments. The genotype will be correlated with respect to clinical microstructural changes for better clinical diagnosis and patient management.
Leber’s hereditary optic neuropathy (LHON):
We intend to do i) Molecular characterisation of nuclear genes involved in LHON; ii)Development of In-Vitro model to understand the involvement of nuclear genes in LHON pathogenesis; and iii) integration of the mito-nuclear genome data with clinical features for improved molecular diagnostics.
In addition, we aim to identify the role of mitochondrial DNA variants, including somatic variations, responsible for Trabecular Meshwork (TM) damage in HTG and NTG, possibly by causing clustered differential expression of cells within TM.
Ongoing Projects
Genetics
- Decoding the unknown genetic etiology to ameliorate the molecular diagnostics of Leber’s Congenital Amaurosis (ICMR-DHR).
- Investigating nuclear gene involvement in a mitochondrial disorder: Leber’s Hereditary Optic Neuropathy (SERB).
- Mitochondrial Association with Transcriptomic Signatures for Trabecular Meshwork Damage in Glaucoma: A tissue-specific omics approach (ICMR).
Genetic testing and counselling of retinoblastoma
India has the highest number of retinoblastoma (RB) patients among the developing countries owing to its increasing population. Our rapid screening strategy identified the spectrum of mutations in more than 90% of patients in a time-efficient and cost-effective manner. Precise risk prediction for siblings and offspring is made possible by genetic counselling.
Novel therapeutics in retinoblastoma
Current therapeutic regimen for retinoblastoma is not a targeted therapy, often leading to adverse effects. A precise molecular therapy will alleviate these side effects and offer better treatment outcomes. We investigate genetic alterations of putative drug targets through targeted NGS approach. The identified copy number alterations in kinases and cell cycle regulators are being tested with FDA approved inhibitors that can serve as promising anti-cancer drugs for the treatment of RB.
Chemoresistance in Retinoblastoma
Chemoresistance remains a major challenge for the clinicians in the treatment of retinoblastoma. We established an in vitro drug resistance model and patient-derived spheroids. Stem cell markers were found to be overexpressed in these systems and currently developing methods to inhibit their growth for overcoming drug resistance.
Molecular characterisation of lymphoma
Ocular Adnexal B Cell Lymphoma (OABL) is the most common orbital neoplasm in adults with disease recurrence, posing a major challenge in treatment. Identifying molecular drivers linked to the lymphoma pathology is crucial for developing effective prognostic markers. Our multi-omic analysis reveals dysregulation of oncogenic signalling pathways, providing insights for newer, effective, and targeted treatment of OABL.
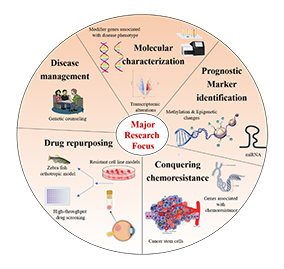
Molecular Biology
- Molecular Characterisation of Ocular Lymphoma for improved Disease Prognosis (Lady Tata Memorial Fellowship)
- Elucidating the role of cancer stem cells in chemoresistant retinoblastoma and their therapeutic implications (ICMR)
- Early Detection of Glaucoma by Genome Analysis (AEF)
Faculty
Research Scholars
- Ms. S. Shiva Sankari
- Ms. K. Saraswathi
- Mr. Sethu Nagarajan
Senior Technician
- Mr. V. Saravanan
Genetic Analyst
- Mr. K. Murugan
Lab Assistant
- Mrs. D. Muthuselvi
- Mrs. T.M. Jayadeepa
Next Generation Sequencing:
The Next Generation Sequencing (NGS) facility at AMRF comprises Illumina Miseq, Covaris, Agilent Tapestation, and a data analysis workstation. DNA is fragmented using the ultrasonication system Covaris. NGS libraries for the metagenomics, targeted exome analysis are prepared using amplicon or probe based chemistries. The prepared libraries are quality checked by Agilent Tapestation. Samples for sequencing are prepared with equimolar concentration and loaded onto the Illumina Miseq system. The generated data are analysed, and reports are generated using the workstation.
The department is equipped with major instruments such as Genetic Analyser, Real-Time PCR, and Bioplex to conduct genomic research.
Services:
- Oculocutaneous and ocular albinism screening
- LCA – RPE65 gene and LHON primary mutations screening for diagnosis
- Genetic counselling for inherited eye disorders
Genetic Testing and Counselling:
Genetic testing of retinoblastoma was carried out, and counselling was done for more than 500 patients and families. This service is now rendered to Aravind-Madurai and other hospitals and multiple centres across the country.
Sequencing:
Sanger sequencing is offered for species identification by 16S, ITS region sequencing. Amplified products of specific genes are sequenced to identify mutations and deletions. This service is utilised by the other labs and academic institutions in and around Madurai. NGS services are provided based on the project requirement.


